|
1/25/2024 0 Comments Rigid protein scaffold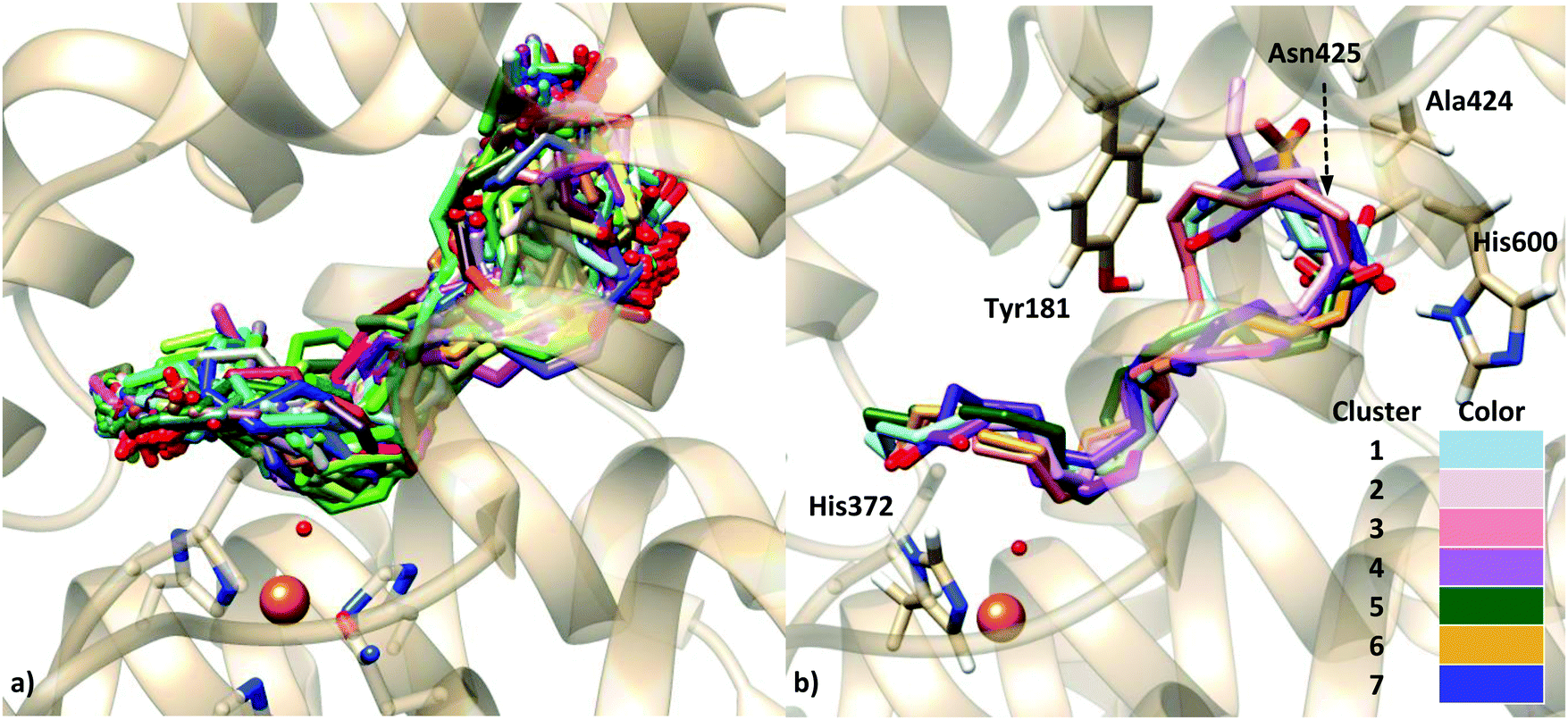
The Legobody approach thus overcomes the current size limitations of cryo-EM = ,Ĭombating the worldwide spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the emergence of new variants demands understanding of the structural basis of the interaction of antibodies with the SARS-CoV-2 receptor-binding domain (RBD). The utility of the method is demonstrated for the KDEL receptor, a 23-kDa membrane protein, resulting in a map at 3.2-Å overall resolution with density sufficient for de novo model building, and for the 22-kDa receptor-binding domain (RBD) of SARS-CoV-2 spike protein, resulting in a map at 3.6-Å resolution that allows analysis of the binding interface to the nanobody. The overall ensemble of ~120 kDa, called Legobody, does not perturb the nanobody–target interaction, is easily recognizable in EM images due to its unique shape, and facilitates particle alignment in cryo-EM image processing. The method is based on the availability of a target-binding nanobody, which is then rigidly attached to two scaffolds: 1) a Fab fragment of an antibody directed against the nanobody and 2) a nanobody-binding protein A fragment fused to maltose binding protein and Fab-binding domains. In this review both the structural features and the practical use of scaffold proteins will be discussed and exemplified.We describe a general method that allows structure determination of small proteins by single-particle cryo-electron microscopy (cryo-EM). The repertoire of protein scaffolds hitherto recruited for combinatorial protein engineering purposes will probably be further expanded in the future, including both additional natural proteins and de novo designed proteins, contributing to the collection of libraries available at present. 
Both biomolecular tools have the potential to complement each other, thus expanding the possibility to find an affinity reagent suitable for a given application. Today, progress in protein library technology allows for the parallel development of immunoglobulin (Ig) as well as scaffold-based affinity reagents. Generally, such artificial receptor proteins should be based on monomeric and small polypeptides that are robust, easily engineered, and efficiently produced in inexpensive prokaryotic expression systems. The quickly growing number of approaches can be classified into three groups: carrier proteins for the display of single variegated loops, scaffolds providing rigid elements of secondary structure, and protein frameworks supporting a group of conformationally variable loops in a fixed spatial arrangement. With respect to broader applicability it should involve a type of polypeptide fold which is observed in differing natural contexts and with distinct biochemical functions, so that it is likely to be adaptable to novel molecular recognition purposes. This concept requires an extraordinary stable protein architecture tolerating multiple substitutions or insertions at the primary structural level. The use of so-called protein scaffolds for the generation of novel binding proteins via combinatorial engineering has recently emerged as a powerful alternative to natural or recombinant antibodies.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
